
Contact Information
417 Roger Adams Lab B-4
600 S Mathews Ave
Urbana, IL 61801
Research Interests
Research Topics
Drug Discovery, Host-Pathogen Interactions, Ion Channels, Membrane Biology, Protein Dynamics, Receptor Biochemistry
Disease Research Interests
Cancer, Infectious Diseases, Metabolic Disorders/Diabetes
Research Description
Protein-Lipid interactions; Membrane Fusion; Lipid Metabolism
Research in the Fratti lab focuses on how the chemical and physical properties of membrane bilayers control the function of proteins. The connection between lipids and protein function are important throughout biology and dysregulation is linked to a wide array of diseases including cancer, diabetes, and Alzheimer’s disease. Therefore, it is important that we understand the fundamentals of protein-lipid interactions so that we can better address their disruption in diseases.
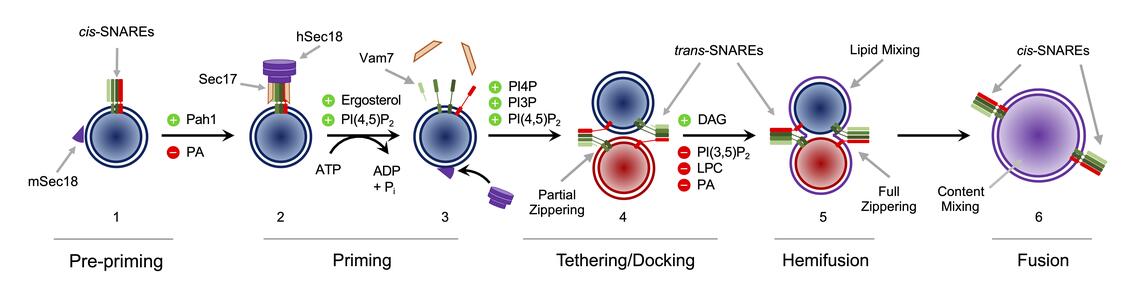
We use lysosomes/vacuoles from the yeast Saccharomyces cerevisiae to reveal how specific lipids inhibit or promote different mechanisms in the process of membrane fusion catalyzed by SNARE proteins. Fusion events are often carried out in highly organized membrane platforms, collectively known as microdomains that are enriched in regulatory lipids that partition from bulk lipids (e.g., phosphatidylcholine) that make up a vesicle. Although low in concentration, regulatory lipids are critical in cellular signaling and the formation of membrane microdomains. The lipids that drive microdomain assembly include phosphatidic acid (PA), diacylglycerol (DAG), phosphoinositides (PI), cholesterol, and sphingolipids. The stoichiometry of these lipids is constantly changing through the action of lipid phosphatases, kinases, and lipases. Lipid modification and remodeling of membranes dramatically changes how lipids interact with proteins and can affect local physical properties such as curvature, fluidity, and bilayer stability. Thus, protein function can be regulated by changes by the lipids in a membrane. Using yeast vacuoles as well as synthetic vesicles we will determine how lipid modification regulates proteins on vesicles as they move through the different stages of the fusion pathway and how dysregulation of these events affects specific genetic and infectious diseases.
Education
B.S. 1992 California State University, Long Beach
Ph.D. 2002 University of Michigan
Postdoc. 2002-2006 Dartmouth Medical School
Additional Campus Affiliations
Highlighted Publications
Representative Publications
Zhang C, Hurst LR, Gokbayrak ZD, Calderin JD, Hrabak MR, Balutowski A, Rivera-Kohr DA, Kazmichuk TDD, Brett CL, Fratti RA. 2024. Sphingolipids regulate Ypt7 function during the tethering stage of vacuole fusion. bioRxiv.
Zhang C, Miner GE, Rivera-Kohr DA, Hrabak M, Balutowski A, Sullivan KD, Guo A, Feng Y, Calderin JD, Fratti RA. 2022. Interdependent transport of Ca2+ and H+ on yeast vacuoles and the role of Phosphatidylinositol 3,5-bisphosphate. J Biol Chem. DOI: https://doi.org/10.1016/j.jbc.2022.102672
Zhang C, Balutowski A, Feng Y, Calderin JD, Fratti RA. 2022. High throughput analysis of vacuolar acidification. Analytical Biochemistry 658:114827.
Sparks RP, Lawless W, Arango AS, Tajkhorshid E, Fratti RA. 2022. Use of Microscale Thermophoresis to Measure Protein-Lipid Interactions. J Vis Exp.(180), e60607, doi:10.3791/60607 .
Fratti RA. 2021. Editorial: Effects of Membrane Lipids on Protein Function. Front Cell Dev Biol. 9:675264.
Sparks RP, Arango AS, Jenkins JL, Guida WC, Tajkhorshid E, Sparks CE, Sparks JD, Fratti RA. 2020. An Allosteric Binding Site on Sortilin Regulates the Trafficking of VLDL, PCSK9 and LDLR in Hepatocytes. Biochemistry. 10.1021/acs.biochem.0c00741.
Miner GE, Sullivan KD, Zhang C, Rivera-Kohr DA, Guo A, Hurst LR, Ellis EC, Starr ML, Jones BC, Fratti RA. 2020. Phosphatidylinositol 3,5-Bisphosphate regulates Ca2+ Transport During Yeast Vacuolar Fusion through the Ca2+ ATPase Pmc1. Traffic. 21:503-517.
Hurst LR, Fratti RA. 2020. Sphingolipids and Lipid Rafts in Yeast Vacuole Fusion and Maturation. Front Cell Dev Biol. 8:539.
Sparks R, Lui A, Bader D, Patel R, Murr M, Guida W, Fratti R, Patel N. 2019. A specific small-molecule inhibitor of Protein Kinase C-delta I activity improves metabolic dysfunction in human adipocytes from obese individuals. J Biol. Chem. 294, 14896-14910.
Sparks RP, Arango AS, Starr ML, Aboff ZL, Hurst LR, Rivera-Kohr DA, Zhang C, Harden KA, Jenkins JL, Guida WC, Tajkhorshid E, Fratti RA. 2019. A small-molecule competitive inhibitor of phosphatidic acid binding by the AAA+ protein NSF/Sec18 blocks the SNARE-priming stage of vacuole fusion. J Biol. Chem. 294, 17168-17185. * Featured Article
Miner GE, Sullivan KD, Zhang C, Hurst LR, Starr ML, Rivera-Kohr DA, Jones BC, Guo A, Fratti RA. 2019. Copper blocks V-ATPase activity and SNARE complex formation to inhibit yeast vacuole fusion. Traffic. 20:841-850.
Starr ML, Sparks RP, Hurst LR, Zhao Z, Arango A, Lihan M, Jenkins JL, Tajkhorshid E, Fratti RA. 2019. Phosphatidic acid induces conformational changes in Sec18 protomers that prevent SNARE priming. J. Biol. Chem. 294:3100-3116.
Starr ML, Fratti RA. 2019. The Participation of Regulatory Lipids in Vacuole Homotypic Fusion. Trends Biochem. Sci. 44:546-554 . * Featured Article
Miner GE, Fratti R. 2019. Real-Time Fluorescence Detection of Calcium Efflux During Vacuolar Membrane Fusion. Methods Mol. Biol. 1860:323-331.
Starr ML, Fratti R. 2019. Determination of Sec18-Lipid Interactions by Liposome-Binding Assay. Methods Mol. Biol. 1860:211-220.
Sparks RP, Jenkins JL, Fratti R. 2019. Use of Surface Plasmon Resonance (SPR) to Determine Binding Affinities and Kinetic Parameters Between Components Important in Fusion Machinery. Methods Mol. Biol. 1860:199-210.
Sparks RP, Fratti R. 2019. Use of Microscale Thermophoresis (MST) to Measure Binding Affinities of Components of the Fusion Machinery. Methods Mol. Biol. 1860:191-198.
Miner GE, Sullivan KD, Guo A, Jones BC, Hurst LR, Ellis EC, Starr ML, Fratti RA. 2019. Phosphatidylinositol 3,5-Bisphosphate Regulates the Transition between trans-SNARE Complex Formation and Vacuole Membrane Fusion. Mol. Biol. Cell. 30:201-208.
Miner GE, Starr ML, Hurst LR, Fratti RA. 2017. Deleting the DAG Kinase Dgk1 Augments Yeast Vacuole Fusion Through Increased Ypt7 Activity and Altered Membrane Fluidity. Traffic. 18:315-329.
Sparks RP, Jenkins LJ, Miner GE, Wangm Y, Guida WC, Sparks CE, Fratti RA, Sparks JD. 2016. Phosphatidylinositol (3,4,5)-trisphosphate binds to sortilin and competes with neurotensin: Implications for very low density lipoprotein binding. Biochem. Biophys. Res. Commun. 479:551-556.
Starr ML, Hurst LR, Fratti RA. 2016. Phosphatidic acid sequesters Sec18p from cis-SNARE complexes to inhibit priming. Traffic. 17:1091-1109.
Sparks RP, Guida WC, Sowden MP, Jenkins JL, Starr ML, Fratti RA, Sparks CE, Sparks JD. 2016. Sortilin facilitates VLDL-B100 secretion by insulin sensitive McArdle RH7777 cells. Biochem. Biophys. Res. Commun. 478:546-552.
Miner G, Starr ML, Hurst LR, Sparks RP Padolina M, Fratti RA. 2016. The Central Polybasic Region of the Soluble SNARE (Soluble N-Ethylmaleimide-sensitive Factor Attachment Protein Receptor) Vam7 Affects Binding to Phosphatidylinositol 3-Phosphate by the PX (Phox Homology) Domain. J. Biol. Chem. 291:17651-17663.
Sasser TL, Fratti RA. 2014. Class C ABC transporters and Saccharomyces cerevisiae vacuole fusion. Cell. Logist. 2014 Jul 3;4(3):e943588.
Lawrence G, Flood B, Brown C, Karunakaran S, Cabrera M, Nordmann M, Ungermann C, Fratti RA. 2014. Dynamic association of the PI3P-interacting Mon1-Ccz1 GEF with vacuoles is controlled through its phosphorylation by the type-1 casein kinase Yck3. Mol. Biol. Cell. 25:1608-1619.
Sasser TL, Lawrence G, Karunakaran S, Brown C, Fratti RA. 2013. The Yeast ATP-binding Cassette (ABC) Transporter Ycf1p Enhances the Recruitment of the Soluble SNARE Vam7p to Vacuoles for Efficient Membrane Fusion. J. Biol. Chem. 288(25):18300-10.
Karunankaran S, Fratti RA. 2013. The Lipid Composition and Physical Properties of the Yeast Vacuole Affect the Hemifusion-Fusion Transition. Traffic. 14(6):650-62.
Sasser TL, Padolina M, Fratti RA. 2012. The yeast vacuolar ABC transporter Ybt1p regulates membrane fusion through Ca2+ transport modulation. Biochem. J. 448(3):365-72.
Karunakaran S, Sasser T, Rajalekshmi S, Fratti RA. 2012. SNAREs, HOPS and regulatory lipids control the dynamics of vacuolar actin during homotypic fusion in S. cerevisiae. J. Cell Sci. 125(Pt 7):1683-92.
Sasser T, Karunakaran S, Qiu Q, Padolina M, Reyes A, Flood B, Smith S, Fratti RA. 2012 The Yeast Lipin 1 Orthologue Pah1p Regulates Vacuole Homeostasis and Membrane Fusion. J. Biol. Chem. 287:2221-2236.
Qiu Q, Fratti RA. 2010. The Na+/H+ exchanger Nhx1p Regulates the Initiation of Saccharomyces cerevisiae Vacuole Fusion. J. Cell Science. 123:3266-3275.
Recent Publications
Sparks, R. P., Lawless, W., Arango, A. S., Tajkhorshid, E., & Fratti, R. A. (2022). Use of Microscale Thermophoresis to Measure Protein-Lipid Interactions. Journal of Visualized Experiments, 2022(180), Article e60607. https://doi.org/10.3791/60607
Zhang, C., Balutowski, A., Feng, Y., Calderin, J. D., & Fratti, R. A. (2022). High throughput analysis of vacuolar acidification. Analytical Biochemistry, 658, Article 114927. https://doi.org/10.1016/j.ab.2022.114927
Zhang, C., Feng, Y., Balutowski, A., Miner, G. E., Rivera-Kohr, D. A., Hrabak, M. R., Sullivan, K. D., Guo, A., Calderin, J. D., & Fratti, R. A. (2022). The interdependent transport of yeast vacuole Ca2+ and H+ and the role of phosphatidylinositol 3,5-bisphosphate. Journal of Biological Chemistry, 298(12), Article 102672. https://doi.org/10.1016/j.jbc.2022.102672
Fratti, R. A. (2021). Editorial: Effects of Membrane Lipids on Protein Function. Frontiers in Cell and Developmental Biology, 9, Article 675264. https://doi.org/10.3389/fcell.2021.675264
Hurst, L. R., & Fratti, R. A. (2020). Lipid Rafts, Sphingolipids, and Ergosterol in Yeast Vacuole Fusion and Maturation. Frontiers in Cell and Developmental Biology, 8, Article 539. https://doi.org/10.3389/fcell.2020.00539