
Contact Information
601 S Goodwin Ave.
Urbana, IL 61801
Research Interests
Research Topics
Archaea, Bioenergetics and Photosynthesis, Drug Discovery, Genetics, Genomics, Regulation of Gene Expression
Research Description
Molecular genetics and biochemistry of methanogenic Archaea; biosynthesis of phosphonate antibiotics; redox cycling of phosphorus by microorganisms
My research program centers on the investigation of two unusual microbial metabolic processes with important biomedical, biotechnological and environmental ramifications. The first project examines the metabolism of reduced phosphorus compounds, with particular emphasis on phosphonic acid antibiotics. The long-term goal of this research is to elucidate the genes and metabolic pathways involved in the biosynthesis of phosphonic acid antibiotics and to explore the molecular diversity of natural products comprising this unusual class of bioactive compounds. We are also interested in the metabolic pathways involved in the catabolism of reduced phosphorus compounds. These studies are also expected to enhance our understanding of phosphorus metabolism, which is central to all living organisms. The second project involves the development and application of genetic techniques for analysis of the methane-producing Archaea. These studies impact a number of critically important to human problems including the production of alternative fuels from biological materials, waste treatment, and global warming.
Microbial metabolism of reduced phosphorus compounds
Unlike the other elements that comprise living matter, it was long believed that phosphorus (P) was redox conservative, existing in the natural world solely in its most oxidized state. Research in my laboratory clearly establishes the capacity of microorganisms to both produce and consume reduced P compounds. Our studies have led to the identification of numerous genes and enzymes required for oxidation of phosphite, hypophosphite and phosphonic acids. Many of these enzymes catalyze unprecedented biochemical reactions. Some have highly useful biotechnological applications, such as the enzyme phosphite dehydrogenase, now being used commercially as a cofactor-regenerating enzyme. Interestingly, many reduced P compounds have potent bioactivities. Investigation of these bioactive compounds is a major area of interest in our group.
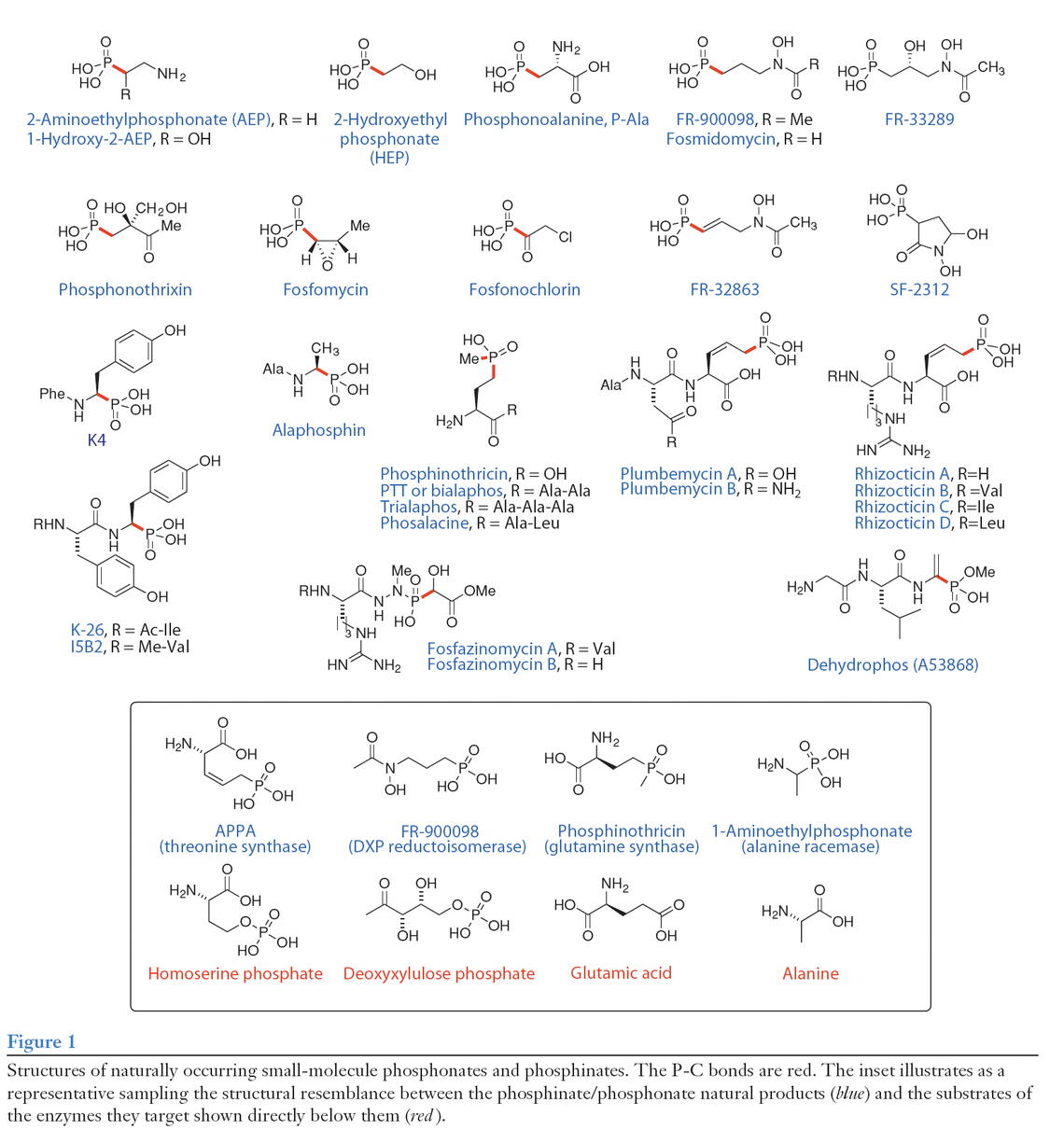
We have focused our research efforts on two particularly promising classes of P compounds, the phosphonic and phosphinic acids. These molecules represent a potent, yet underexploited, group of compounds with great promise in the treatment of human disease. Numerous, structurally distinct, phosphonates are produced in nature and many have useful bioactive properties, Figure 1. Among these are the bioactive compounds phosphinothricin tripeptide (PTT), phosphonothrixin, fosfomycin, plumbemycin, A53868, FR-900098 and fosmidomycin. The latter two are remarkable in that they are highly effective in the treatment of malaria (including multi-drug resistant strains) and various bacterial infections. Fosfomycin, clinically used under the name Monurol®, is effective against a variety of infections and is the only FDA approved drug for treatment of acute cystitis during pregnancy. Phosphinothricin-tripeptide and phosphonothrixin, are excellent herbicides and various formulations of the former are widely used in agriculture, with annual sales in excess of $200 million. Additional medical applications can be found in other naturally produced phosphonates. For example, the phosphonate compound designated K-26 is a potent angiotensin converting enzyme (ACE) inhibitor and shows promise in the treatment of high blood pressure. Importantly, the targets of phosphonate antibiotics vary substantially, allowing their use in the treatment of a wide variety of health conditions. Further, the potent bioactivity of some phosphonates allows their use as fungicides, herbicides and pesticides in agriculture.
In ongoing studies conducted at the Institute for Genomic Biology our group has cloned, sequenced and characterized the gene needed for synthesis of many of the phosphonate compounds shown in Figure 1. These studies have led to the development of engineered strains that overproduce the antibiotics in question, thus lowering the costs associated with their production. Moreover, we have uncovered a wealth of interesting and unprecedented metabolism and biochemistry during our characterization of the antibiotic biosynthetic pathways. Perhaps most importantly, we have shown that genes for phosphonate biosynthesis are quite common in antibiotic-producing microorganisms. Current research is aimed at characterizing previously known and, as yet uncharacterized, phosphonate compounds with the goal of discovering and developing novel therapeutic agents.
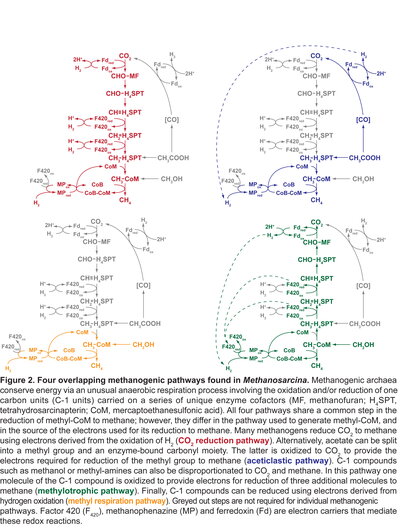
Genetic analysis of methanogenic Archaea Methanogenesis plays an essential role in the biosphere. Each year an estimated 5 x 1014 g of biologically produced methane is released into the atmosphere. Depending on your viewpoint, this represents a staggering untapped renewable energy source and/or a frightening contribution to global warming. Significantly, this value grossly underestimates the flux of carbon via the process because microorganisms consume the majority (50-75%) of methane produced before it can escape into the atmosphere. Therefore, the actual rate of methanogenesis is 2-4 fold higher than the typically quoted value. Moreover, vast amounts of biologically produced methane remain trapped at the sea floor in the form of methane-hydrate: by some estimates a quantity twice that of all other known fossil fuel reserves combined. While these huge numbers clearly establish the essential role of methanogenesis in the global carbon cycle, they fail to capture the full impact of methanogens on biological nutrient cycles and global climate. Methanogenic microorganisms play a keystone role in anaerobic environments whenever more favorable electron acceptors such as Fe(III) or sulfate are absent. In such environments, a complex microbial community ferments biomass to a mixture of acetate, formate, hydrogen and CO2. These compounds, in turn, serve as growth substrates for methanogens. Interestingly, the two processes are inextricably linked. By consuming the fermentation products as rapidly as they are produced (keeping the concentration of these compounds very low), the methanogens render an otherwise endergonic process thermodynamically favorable. Thus, the fermenting microbes are incapable of consuming their growth substrates in the absence of methanogens, whereas the methanogens are completely dependent on the fermenting microbes to produce the substrates they require for growth. This process, known as “syntrophy” (literally translated as “eating together”), is the basis of the anaerobic food chain in most freshwater ecosystems. As a result, anaerobic decomposition of typical biomass produces methane and carbon dioxide in a one to one ratio. Thus, the fraction of the global carbon cycle that is dependent on methanogenesis is actually twice the amount of methane produced. Accordingly, the total amount of carbon turnover that is dependent on methanogens could be as high as 5 x 1015 g per year: roughly 4% of annual primary productivity of the biosphere! The potential impact of methanogens on a number of human issues cannot be ignored. The vast amounts of methane produced in the biosphere have been touted as a renewable, carbon neutral energy source. Methanogenesis is also critically important in agriculture due to the role of methanogenic organisms in ruminant nutrition and waterlogged fields (e.g. rice paddies) and it is a required step in the processing of waste in sewage treatment facilities and landfills. It is estimated that roughly half of biological methane production can be directly attributable to these human activities. As a result, the level of methane in the atmosphere has increased dramatically since the industrial revolution, doubling in the last 1-200 years. This increase carries particular significance to the global climate because methane is a highly potent greenhouse gas, ca. 25-fold more effective than CO2. Although present at much lower concentrations in the atmosphere than CO2, methane currently accounts for ca. 20% of the radiative forcing of all greenhouse gasses. Taken together, these issues provide a compelling rationale for further study of methanogenic microorganisms. My laboratory uses a combination of genetic, biochemical and molecular methods to study the methanogenic archaeon Methanosarcina. Over the past decade we have developed a powerful suite of genetic methods that can be used in many Methanosarcina species. These include: plasmid shuttle vectors, high efficiency transformation, directed gene replacement, in vivo transposon mutagenesis, multiple selectable markers, reporter gene technologies, and an anaerobic incubator for large-scale growth of methanoarchaea on solid media. Genetic manipulation of Methanosarcina species is simple and reliable and these methods have proven invaluable for studying the metabolism and physiology of multiple Methanosarcina species.
Education
B.S. (Anthropology), University of Illinois, 1981
B.S. (Microbiology), University of Illinois, 1984
Ph.D. (Microbiology), Purdue University, 1991
Postdoctoral (Microbiology), Purdue University, 1991-1993
Postdoctoral (Microbiology), University of Illinois, 1993-1997
Awards and Honors
Fellow in the American Association for the Advancement of Science (AAAS), 2016
Fellow in the American Academy of Microbiology, 2010
Additional Campus Affiliations
External Links
Highlighted Publications
Representative Publications
1. Parkinson, E.I., A. Erb, A.C. Eliot, K-.S. Ju and W.W. Metcalf. 2019. Fosmidomycin biosynthesis diverges from related phosphonate natural products. Nat. Chem. Biol. in press.
2. Mand, T.D. and W.W. Metcalf. 2019. Energy conservation and hydrogenase function in methanogenic archaea with a focus on the genus Methanosarcina. Microbiol. Mol. Biol. R. in press.
3. Nayak, D.D. and W.W. Metcalf. 2019. Methylamine-specific methyltransferase paralogs in Methanosarcina are functionally distinct despite frequent gene conversion. ISME J. in press. PMID: 31053830.
4. Fu, H., M.N. Goettge and W.W. Metcalf. 2019. Biochemical characterization of the methylmercaptopropionate:Cob(I)alamin methyltransferase from Methanosarcina acetivorans. J. Bacteriol. J Bacteriol. 201: pii: e00130-19. PMID: 30936368.
5. Petronikolou, N., M.A. Ortega, S.A. Borisova, S.K. Nair and W.W. Metcalf. 2019. Molecular basis of Bacillus subtilis ATCC 6633 self-resistance to the phosphono-oligopeptide antibiotic rhizocticin. ACS Chem Biol. 14:742-750. PMID: 30830751.
6. Nayak, D.D. and W.W. Metcalf. 2018. Genetic techniques for studies of methyl-coenzyme M reductase from Methanosarcina acetivorans C2A. Methods Enzymol. 613:325-347. PMID: 30509472.
7. Mand, T.D., G. Kulkarni and W.W. Metcalf. 2018. Genetic, biochemical, and molecular characterization of Methanosarcina barkeri mutants lacking three distinct classes of hydrogenase. J. Bacteriol. 200: e00342-18. PMID: 30012731
8. Kulkarni, G., T.D. Mand and W.W. Metcalf. 2018. Energy Conservation via Hydrogen Cycling in the Methanogenic Archaeon Methanosarcina barkeri. mBio 9:e01256-18. PMID: 29970471
9. Goettge, M.N., Cioni, J.P., Ju, K-.S., Pallitsch, K. and W.W. Metcalf. 2018. PcxL and HpxL are flavin-dependent, oxime-forming N-oxidases in phosphonocystoximic acid biosynthesis in Streptomyces. J. Biol. Chem. 293:6859-6868. PMID: 29540479.
10. Parkinson, E.I., Tryon, J.H., Goering, A.W., Ju, K-.S., McClure, R.A., Kemball, J.D., Zhukovsky S., Labeda, D.P., Thomson, R.J., Kelleher, N.L., and W.W. Metcalf. 2018. Discovery of the Tyrobetaine Natural Products and Their Biosynthetic Gene Cluster via Metabologenomics. ACS Chem. Biol. 13:1029-1037. PMID: 29510029.
12. Nayak, D.D., Mahanta, N., Mitchell, D.A. and W.W. Metcalf. 2017. Post-translational thioamidation of methyl-coenzyme M reductase, a key enzyme in methanogenic and methanotrophic Archaea. Elife 6: e29218. PMID: 28880150
13. Nayak, D.D. and W.W. Metcalf. 2017. Cas9-mediated genome editing in the methanogenic archaeon Methanosarcina acetivorans. Proc. Natl. Acad. Sci. USA 114:2976-2981. PMID: 28265068, PMC5358397
Recent Publications
Zimmermann, A., Xia, S. N., Moschny, J., Gomez-Escribano, J. P., Boldt, J., Nübel, U., Nouioui, I., Krause, J., Irle, M. K., Metcalf, W. W., Hughes, C. C., & Mast, Y. (2026). Expanding the actinomycetes landscape for phosphonate natural products through genome mining. RSC Chemical Biology, 7(2), 298-312. https://doi.org/10.1039/d5cb00254k
Bonitatibus, S. C., Pham, R. V., Weitz, A. C., Lopéz-Muñoz, M. M., Li, B., Metcalf, W. W., Nair, S. K., & Elliott, S. J. (2025). The redox landscape of pyruvate:ferredoxin oxidoreductases reveals often conserved Fe–S cluster potentials. Journal of Biological Chemistry, 301(8), Article 110380. https://doi.org/10.1016/j.jbc.2025.110380
Metcalf, W. W., Polidore, A., & Kloeber, J. (2025). Phosphonate products and methods. (U.S. Patent No. 12264172).
Rodriguez Carrero, R. J., Lloyd, C. T., Borkar, J., Nath, S., Mirica, L. M., Nair, S., Booker, S. J., & Metcalf, W. (2025). Genetic and biochemical characterization of a radical SAM enzyme required for post-translational glutamine methylation of methyl-coenzyme M reductase. mBio, 16(2). https://doi.org/10.1128/mbio.03546-24
Weinberger, V., Mohammadzadeh, R., Blohs, M., Kalt, K., Mahnert, A., Moser, S., Cecovini, M., Mertelj, P., Zurabishvili, T., Arora, B., Wolf, J., Shinde, T., Madl, T., Habisch, H., Kolb, D., Pernitsch, D., Hingerl, K., Metcalf, W., & Moissleichinger, C. (2025). Expanding the cultivable human archaeome: Methanobrevibacter intestini sp. nov. and strain Methanobrevibacter smithii ‘GRAZ-2’ from human faeces. International Journal of Systematic and Evolutionary Microbiology, 75(4), Article 006751. https://doi.org/10.1099/ijsem.0.006751




