The task: calculate every movement of every human in New York City and, based on those data, create a picture-perfect map of the city's skyline. Seem impossible?
And yet, this resembles what two School of Molecular & Cellular Biology scientists recently accomplished while investigating E. coli membrane transport proteins — a system with billions of more variables and on an atomic scale.
In new research, Dr. Emad Tajkhorshid, J. Woodland Hastings Endowed Chair and Professor of Biochemistry, PhD candidates Yupeng Li, Carlos Cuellar Rodriguez, Dr. Hale Şiir Hasdemir (PhD, ’25, biophysics), and their colleagues mapped out a new family of membrane transport protein responsible for phospholipid transport. Their findings may offer new insights into developing novel therapies that combat antibiotic resistance.
The article, “LetA defines a structurally distinct transporter family,” was published in Nature earlier this year.
Escherichia coli, long a model organism for biochemical researchers, has an outer cell membrane that is dense and thick. The heavily fortified membrane “maintains the cell’s integrity so that it can protect itself from other toxic molecules, like antibiotics,” Li said.
When an antibiotic encounters a cellular membrane, it’s met with a wall of lipids.
How E. coli transports these tricky, amphiphilic lipids from inside the cell, across its inner membrane, through the long periplasmic space, and integrates them into the outer membrane isn’t fully understood. Although several components involved in this process have been identified, the detailed molecular mechanisms behind phospholipid transport are still being investigated.
To find out how the lipid transit system worked, the team started by looking at E. coli’s transport proteins. These proteins vary by organism and cell type, but all act as any cell’s entry and exit points. “These proteins are responsible for moving molecules from one side of the membrane to the other,” Tajkhorshid said.
There were a few hurdles towards determining how the transport protein functioned. First, E. coli’s outer membrane, where phospholipids are needed, has no energy source. Its job is to protect, not to move. “It cannot do any of the work transporting lipids by itself,” Tajkhorshid said. Also, this process is highly dynamic and occurs on an atomic scale, undetectable to the human eye and difficult to capture even with advanced experimental techniques.
The only way to get a picture of what was happening was with a “molecular dynamic simulation.” Simply put, if one can calculate the placement and movement of every single atom in a system for a fraction of a second, then their behavior can be simulated, and what’s exactly pushing the lipid through might be revealed.

“In these simulations, you’re essentially looking at timestamps of 10-15 seconds,” Tajkhorshid said. “That means if you even want to look for a microsecond, you have to calculate all of the forces between all the atoms in the system, which is usually on the order of billions of times. Then, we have to translate these dynamics into the function of a protein system.”
Those systems usually follow a similar pattern. “For proteins that transport small molecules, usually they have structures like this,” Li said, making a “V” shape with his arms. “In this usual mechanism the transport proteins can change conformations to the ‘V’ or its reverse so molecules can be transported in or out.”
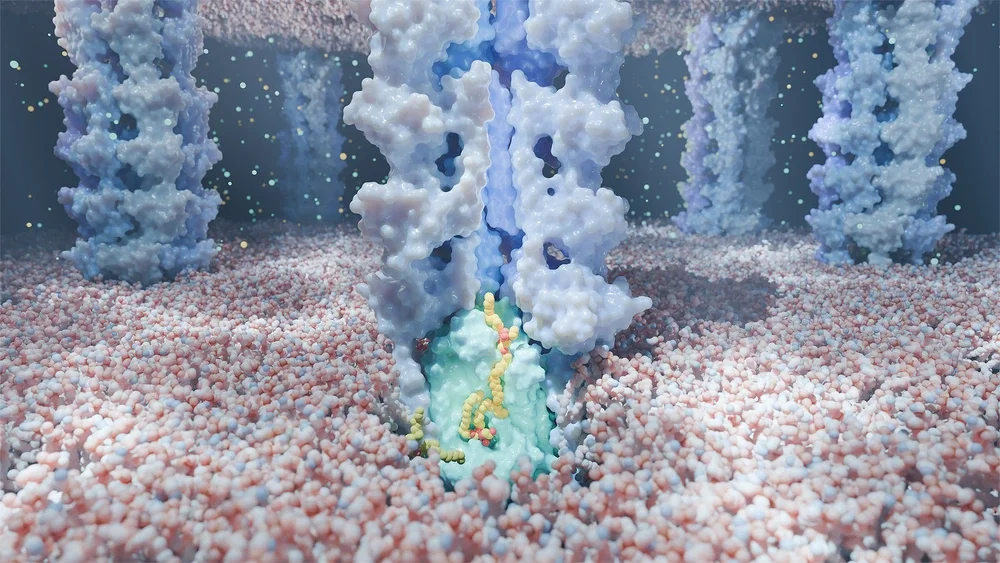
However, the mechanism that Tajkhorshid and Li’s data revealed was something scientists had never seen before. E. coli’s phospholipid transport system, they found, is made up of two components — LetA and LetB. With a lipid as the cargo, these two proteins work similar to a tugboat (LetA) guiding a ship into a canal (LetB).
“Let A recruits the lipids from the inner membrane, lifts them, and then pushes them in to LetB,” Tajkhorshid said. “LetB is a tunnel that allows lipids to diffuse inside it, so their water-averse portions can be shielded during transport and the entire lipids can be incorporated into the outer membrane.”
LetA working with LetB also solved the mystery behind how the outer membrane has no energy but is still able to incorporate lipids. A canal doesn’t need any energy if a tugboat does all the work.
After their data revealed the LetA-LetB mechanism, Tajkhorshid and Li’s coauthors were able to deduce that E. coli’s transport proteins were evolutionarily related to specific non-transporter proteins of humans.
“If we can understand this mechanism — how Gram-negative bacteria can transport lipids all the way from the inner membrane to the outer membrane — then we can probably develop novel therapeutics that target this system and combat antibiotic resistance,” Li said.
Dr. Tajkhorshid and Li would like to thank the biophysics program, the National Center for Supercomputing Applications at the University of Illinois, and the National Institutes for Health for supporting the study and for their contributions to running the simulations which revealed the LetA-LetB mechanism.
