
Contact Information
116A Morrill Hall
505 S. Goodwin
Urbana, IL 61801
Biography
Stephen G. Sligar received his Ph.D. in Physics from the University of Illinois in 1975. Dr. Sligar served on the faculty in the Department of Molecular Biophysics and Biochemistry at Yale University and returned to the University of Illinois in 1982 where he was the I. C. Gunsalus Professor of Biochemistry. He now holds the University of Illinois Swanlund Endowed Chair and is Director of the School of Molecular and Cellular Biology. He is also a faculty member in the Department of Chemistry, the Center for Biophysics and Computational Biology and the College of Medicine. Dr. Sligar holds affiliate appointments in the Beckman Institute for Advanced Science and Technology, the Institute for Genomic Biology and The Micro and Nano Technology Laboratory on the Illinois campus. He is a Fellow of the Biophysical Society and the American Association for the Advancement of Science. Awards include a Fulbright Research Scholarship, Senior Fellowship from the Japan Society for the Promotion of Science, an NIH Merit Award and the Bert L. and Kuggie Vallee Visiting Professorship in Inorganic Chemistry at Oxford where he was a Fellow of Queens College. He is also a Fellow in the Jerome Karle Nobel Laureate World Innovation Foundation. Dr. Sligar's research is supported by grants from the National Science Foundation, the National Institutes of Health and the Human Frontiers Program. Research centers on understanding the structure and mechanistic function of metalloenzymes, membrane bound receptors and transporters as well as investigations in blood coagulation and amyloid proteins and their corresponding human disease states.
Research Interests
physical and chemical mechanisms of oxygenase catalysis; self-assembled nanoscale complexes; development of therapeutics for human disease targets
Research Description
Structural and functional characterization of macromolecular assemblies. Hormone biosynthetic cancer targets. Mechanisms of drug metabolism. Cancer signaling through the Ras pathways.
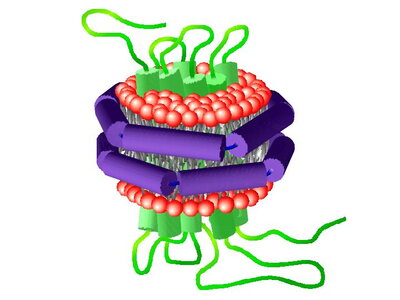
A major goal of our laboratory is to understand the mechanisms involved in cancer signaling. In many cases, the relevant signaling complexes assemble on a membrane surface, yet the role of this critical component has not been investigated. We use the Nanodics system and extensive long-term molecular dynamics simulations to define the role of the membrane in determining the structure and interactions of the oncogenic protein KRas4b with the membrane surface and with its effector proteins. The Rat sarcoma (Ras) family of small membrane-associated GTPases are essential molecules involved in a signal transduction cascade that regulate, among other cellular properties, survival and proliferation. Central to Ras-mediated signal transduction is proper transport from the endoplasmic reticulum and subsequent stable association with the lipid bilayer of the plasma membrane where, once activated, Ras recruits downstream effector proteins leading to their subsequent activation.
A second major theme is the cytochrome P450 monosygenases involved in human drug metabolism and steroid hormone biosynthesis. The cytochromes P450 are members of a class of enzymes known oxygenases, as the incorporate an oxygen atom from atmospheric dioxygen into a substrate molecule. Our major efforts are focused on isolating and characterizing the reactive intermediates of the P450 catalytic cycle, the modes of molecular recognition of enzyme for its substrate and the details of inter-protein electron and proton transfer. For the drug metabolizing P450s, we seek to understand the molecular mechanisms of drug-drug interaction and the role of protein dynamics in linking the allosteric and catalytic sites of the enzymes. In the case of steroid biosynthesis, we are identifying the specific catalytic intermediate involved in catalysis with a goal of developing novel inhibitors of these enzymes that are central targets in the treatment of hormone dependent cancers.
Education
B.S. 1970 Drexel University
Ph.D. 1975 University of Illinois, U-C
Postdoc. 1975-76 University of Illinois, U-C
Awards and Honors
Member, National Academy of Sciences
Christian Anfinsen Award, Protein Society
Sober Award, American Society of Biochemistry and Molecular Biology
Fellow, Biophysical Society
NSF Human Frontiers Science Program Research Award
World Innovation Foundation Fellow
NIH Merit Award
Japan Society for the Promotion of Sciences Senior Fellow
Bert L. and N. Kuggie Vallee Visiting Professor, Oxford University
Fellow, AAAS
Additional Campus Affiliations
Maybelle Leland Swanlund Endowed Chair Emeritus, Biochemistry
Professor Emeritus, Biochemistry
Research Professor, Biochemistry
Professor Emeritus, Chemistry
External Links
Highlighted Publications
Representative Publications
Over 400 Publications; Google Scholar h-index = 111; 46,630 total citations. Selected recent publications:
Liu, Y., Grinkova, Y., Gregory, M.C., Denisov, I.G. and Sligar, S.G. (2021) “Mechanism of the Clinically Relevant E305G Mutation in Human P450 CYP17A1” Biochemistry 60, 3262-3271
Denisov, I.G., Grinkova, Y.V., Camp, T., McLean, M.A. and Sligar, S.G. (2021) “Midazolam as a Probe for Drug-Drug Interactions Mediated by CYP3A4: Homotropic Allosteric Mechanism of Site-Specific Hydroxylation” Biochemistry 60, 1670-1681.
Liu, Y., Denisov, I.G., Sligar, S.G. and Kincaid, J.R. (2021) “Substrate-specific Allosteric Effects on the Enhancement of CYP17A1 Lyase Efficiency by Cytochrome b5” J. American Chemical Society 143, 3729-3733.
Sligar, S.G. and Denisov, I.G. (2020) “Nanodiscs: A Toolkit for Membrane Protein Science” Protein Science 30, 297-315.
Liu, Y., Denisov, I.G., Grinkova, Y.V., Sligar, S.G. and Kincaid, J.R. (2020) “P450 CYP17A1 Variant with a Disordered Proton Shuttle Assembly Retains Peroxo-Mediated Lyase Efficiency” Chemistry 26, 16846-16852.
Camp, T. and Sligar, S.G. (2020) “Nanodisc Self-Assembly is Thermodynamically Reversible and Controllable” Soft Matter 16, 5615-5623.
Gorfe, A. and Sligar, S.G. (2020) “Membrane Bound Ras as a Conformational Clock” Biophysical Journal 118, 1-3.
Camp, T., Mehta, K., Sligar, S.G. and Zhang, K. (2020) "Molecular Orientation Determination in Nanodiscs at the Single Molecule Level" Analytical Chemistry 92, 2229-2236.
McLean, M.A., Stephen, A.G. and Sligar, S.G. (2019) "PIP2 Influences the Conformational Dynamics of Membrane Bound KRAS4b" Biochemistry 58, 3537-3545.
Denisov, I.G., Grinkova, Y., Nandigrami, P., Shekhar, M., Tajkhorshid, E. and Sligar, S.G. (2019) "Allosteric Interactions in Human Cytochrome P450 CYP3A4: The Role of Phenylalanine 213" Biochemistry 58, 1411-1422.
Wong, P., Li, L., Chea, J., Poku, E., Ebner, T. Bowkes, N., Wong, J. Y. C., Yazaki, P. J., Sligar, S. G., and Shively, J. E. (2020) “Antibody targeted PET Imaging of 64Cu-DOTA-Anti-CEA PEGylated Lipid Nanodiscs in CEA positive tumors” Bioconjugate Chemistry 31, 743-753.
Mustafa, G., Nandekar, P.P., Camp, T.J., Bruce, N.J., Gregory, M.C., Sligar, S.G. and Wade, R.C. (2019) “Influence of Transmembrane Helix Mutations on Cytochrome P450-Membrane Interactions and Function” Biophysical Journal 116, 1-14.
Recent Publications
Denisov, I. G., Grinkova, Y. V., & Sligar, S. G. (2026). Lipid Dependence of CYP3A4 Activity in Nanodiscs. Biology, 15(2), Article 156. https://doi.org/10.3390/biology15020156
Denisov, I. G., Liu, Y., Mak, P. J., & Sligar, S. G. (2026). Mechanistic and spectroscopic characterization of human CYP17A1 in Nanodiscs. Journal of Inorganic Biochemistry, 277, Article 113211. https://doi.org/10.1016/j.jinorgbio.2025.113211
Reyes-Ordoñez, A., Shree, S., Sligar, S. G., & Chen, J. (2026). Protocol for Nanodisc single-molecule pull-down assay to detect protein-lipid interactions. STAR Protocols, 7(1), Article 104377. https://doi.org/10.1016/j.xpro.2026.104377
Reyes-Ordoñez, A., Shree, S., Singh, N., Sligar, S. G., & Chen, J. (2025). Nanodisc single-molecule pulldown to study lipid-protein interactions. Journal of Lipid Research, 66(7), Article 100846. https://doi.org/10.1016/j.jlr.2025.100846
Denisov, I. G., & Sligar, S. G. (2024). Nanodiscs for the study of membrane proteins. Current Opinion in Structural Biology, 87, Article 102844. https://doi.org/10.1016/j.sbi.2024.102844

